Cohorts that respect consent.
Atlas-style cohort builders fetch by criteria and ignore consent context. Forge is built differently: every query is consent-scoped, every access is HAVEN-audited. The cohort you ship is the cohort you can defend — running today on a simulated MIMIC-IV cohort.
Three pains every clinical researcher knows.
Your cohort is a compliance gamble
Atlas, i2b2, and EDW exports fetch by criteria and ignore consent context. Your published cohort may include patients who never consented to that purpose. You won’t know until someone asks.
No audit chain to hand IRB
When IRB or a journal asks who consented to what, you compile from spreadsheets, emails, and EHR audit logs. There is no single, signed chain you can hand over. There is no chain at all.
Multi-site cohorts die at the paperwork
Cross-institutional data sharing requires BAA, DUA, IRB reciprocity per pair of institutions. Months of legal before a single query runs. Most multi-site studies fail here, not at the science.
Forge as the cohort builder. HAVEN underneath. PSDL for cross-site reproducibility when you need it.
Forge
HAVEN
Prometheno
PSDL family
Same scenario. Two simulated sites. Byte-identical hashes.
An AKI Detection scenario, authored once in psdl-inspector, compiled to a Certified Bundle, executed against MIMIC-IV (Site A) and a simulated dataset (Site B). The output hashes match. That is what reproducibility looks like at the protocol layer.
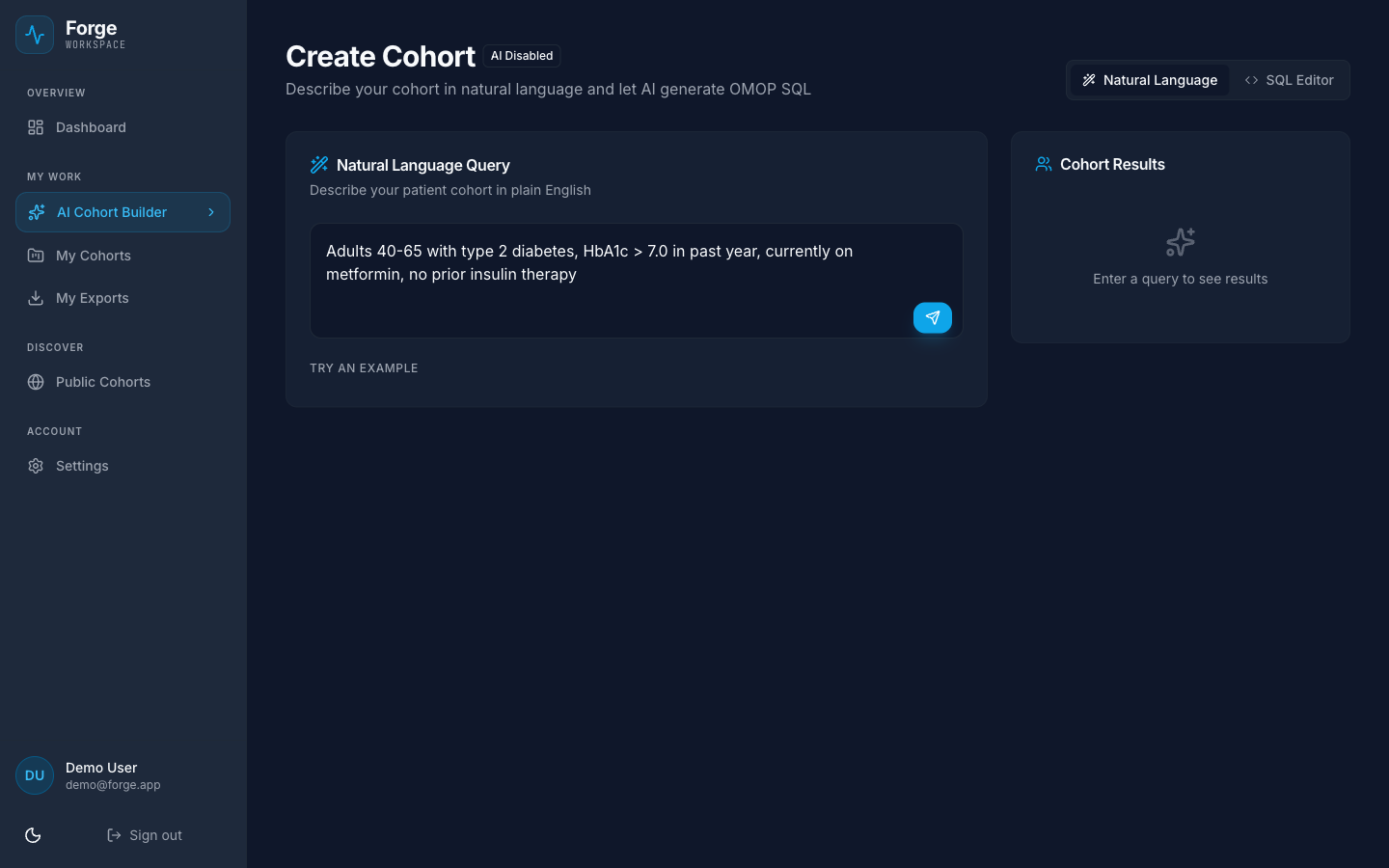
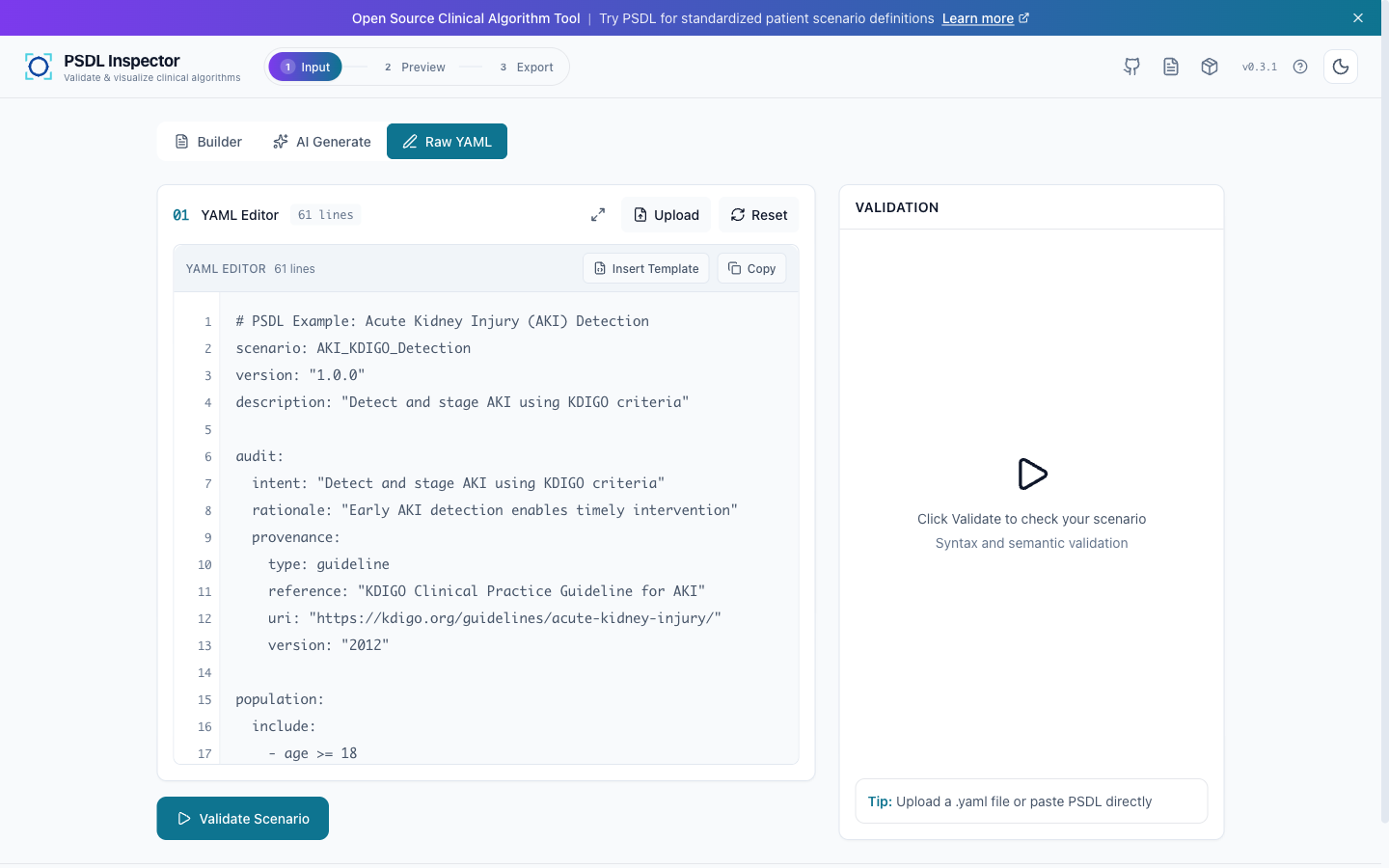
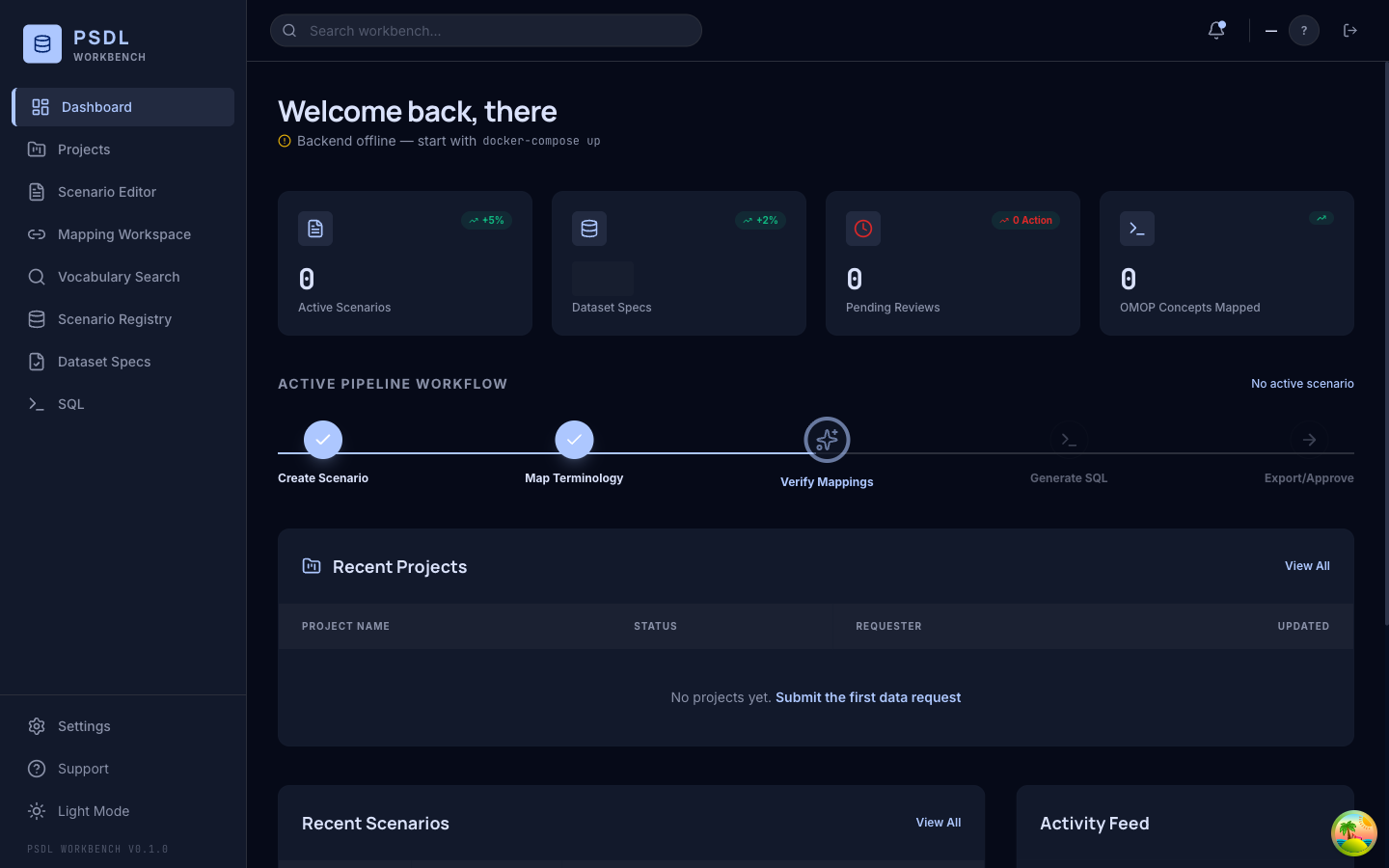
Author
AKI scenario in psdl-inspector with KDIGO criteria.
scenario: AKI_Detection audit: intent: ... rationale: KDIGO 2012 signals: { Cr: ... } trends: { delta_6h: ... } logic: { aki_risk: ... }Certify
Compile to a Certified Bundle. SHA-256 binds everything.
{ "bundle_version": "1.1", "checksum": "sha256:abc123...", "scenario": { ... }, "terminology_anchors": { "creatinine": { "concept_id": 3016723, "vocabulary_id": "LOINC" } } }Run · Site A
Execute on MIMIC-IV via Prometheno. Hash recorded in HAVEN audit chain.
$ psdl run bundle.json --site mimic-iv Triggered patients: 47 Result IR hash: 0x9799d065d19f...e583 HAVEN audit entry: #1247Run · Site B
Same bundle, different dataset. Hash identical. Reproducibility, proven.
$ psdl run bundle.json --site sim-b Triggered patients: 31 Result IR hash: 0x9799d065d19f...e583 HAVEN audit entry: #889 ✓ Hashes matchThe MIMIC-IV Colab notebook runs end-to-end in your browser. No install.
Hashes shown are illustrative. Actual hashes match across runs of identical inputs.
When this works at scale.
Cross-institutional cohorts in hours, not months. Multi-site reproducibility as default. IRB documentation auto-generated from the same source as execution. Researchers spend their time on science instead of data acquisition.
We sell access tiers — academic discount, industry rates — to consented cohorts and to the platform that runs them. The protocols underneath stay open.
The work is open. Pick your level.
MIMIC-IV Colab notebook. No install. End-to-end PSDL on real ICU data.
PSDL on your dataset. Adapter pattern documented. Tell us where the spec fails.
Adversarial spec review. Ambiguities, edge cases, contradictions.
Run a clinical research program? Test cross-institutional cohorts on your data.